Journal of Oil Palm Research 33 (2) June 2021, p. 215-226
SOIL FUNGAL COMPOSITION AND DIVERSITY IN OIL PALM PLANTATION AT SUNGAI ASAP, SARAWAK, MALAYSIA
DOI: https://doi.org/10.21894/jopr.2021.0014
Received: 12 July 2017 Accepted: 29 August 2019 Published Online: 15 April 2021
A study was carried out to compare the soil fungal composition and diversity in oil palm plantation with the adjacent secondary forest at Sungai Asap, Sarawak, Malaysia using denaturing gradient gel electrophoresis (DGGE). Findings revealed that soil fungal composition in the oil palm plantation was significantly different compared with undisturbed secondary forest (SF1) and disturbed secondary forest (SF2) by forming three main clusters as shown in the principal component analysis (PCA). Furthermore, DGGE profile revealed that the banding pattern of oil palm planted area (OPPA) remained diverse throughout the assessment years. However, DGGE banding patterns of SF1 and SF2 showed similarities as noted in the dry season (June 2011) whereby both clusters were overlapped. The composition of soil fungal community in oil palm plantation was dominated by fungi from the phylum Ascomycota whereas secondary forests were dominated by fungi from Basidiomycota. Biodiversity indices showed that the soil fungal diversity in SF1 was higher than SF2 and OPPA. It was postulated that there was possible interactions between soil fungal communities with plantation activities thereby soil fungal communities were shifted in order to acclimatise with new environments in the plantation.
KEYWORDS:Oil palm (Elaeis guineensis) is cultivated commercially in Indonesia and Malaysia as a main source of cooking oil. Oil palm plantation is a highly important industry because of significant contribution to the Malaysian economy. This sector occupies 5%-6% of Malaysia’s gross domestic product (GDP) (Malaysian Palm Oil Council, 2015). Also, palm oil industry is very successful to eradicate poverty in Malaysia (Palm Oil World, 2011). Sarawak is one of the largest palm oil producers in Malaysia, and has registered a total of 1.44 million hectares of oil palm planted areas (MPOB, 2016). Due to the widespread of plantation, environmental issues such as deforestation and the loss of biodiversity have arisen (Fitzherbert et al., 2008). Reduced biodiversity may have other negative impacts on ecosystem functions, especially on the soil microorganisms’ which play vital roles in maintaining soil ecosystem.
Although, the oil palm industry is highly significant to the Malaysian economy but the establishment of vast monoculture system without proper legal control has created a number of environmental impacts. The two most serious are large-scale forest conversion and loss of critical habitat for endangered species. Other associated issues are soil erosion, air, soil and water pollutions as well as climate change. Furthermore, new plantations can also create social conflicts if the rights and livelihoods of local communities are ignored. By studying composition of soil microorganisms their diversity and their roles in ecological functions may help in improving quality of soil and ecosystem health. The outputs of the research findings can be used to develop specific programme and strategy in order to conserve and sustain our mother nature for the future generation.
Soil is one of the most diverse terrestrial habitats that support life forms from microorganisms to invertebrates, flora and fauna. Microorganisms are the largest and the most diverse biotic group in soil, with an estimated one billion microorganisms per one gram of soil (McCauley et al., 2005). They are bacteria, fungi, actinomycetes, algae and protozoa. They mediate major processes occurring in the soil including decomposition activity, nutrient and energy cycling. Several difficulties involved in studying soil microbial community were reported. Besides, variation in environmental factors in each of the sampling sites may cause difficulties in comparing the results.
Meanwhile, advancement in research techniques especially in molecular biology has opened up a new dimension in studying environmental functions. Molecular approaches like polymerase chain reaction (PCR) techniques and the fingerprinting techniques, such as denaturing gradient gel electrophoresis (DGGE), temporal temperature gradient gel electrophoresis (TTGE) and amplified recombinant deoxyribonucleic acid (rDNA) restriction analysis (ARDRA), terminal restriction fragment length polymorphism (T-RFLP) are widely used to study the diverse community members in any given environment. These molecular methods do not require the extraction of individual (DNA) or to culture the organisms, yet can be targeted at a very broad or narrow range of organisms present in the community (Torsvik and Øvreås, 2002).
In soil ecosystems, fungi play fundamental roles during decomposition of plant residues which promotes nutrient cycling and stimulate plant growth. In addition, soil fungi along with other soil microbes are significant in conserving soil productivity and are sensitive to the small changes in soil environment. Hence, studying soil fungal community is a good way to determine soil health and quality. In other words, soil fungal communities have the potential role as a biological indicator for the soil quality and health. Thus, the aims of this study were to determine the short-term impacts on soil fungal composition and diversity in oil palm plantation using PCR-DGGE method.
Soil Sampling
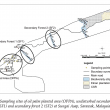
Soil samples were collected from the Malaysian Palm Oil Board (MPOB) Research Station at Sungai Asap, Sarawak, Malaysia. Samplings were carried out at a six-month interval (December 2010, June 2011 and December 2011). There were three sampling sites, the undisturbed secondary forest (SF1), disturbed secondary forest (SF2) and oil palm planted area (OPPA). The OPPA was 106.4 ha. SF1 and SF2 are two forests nearby, where SF1 was 6 km and SF2 was 9.7 km from OPPA. Size of the sampling sites for both SF1 and SF2 were approximately 30 ha. During the study time, the oil palms were three years old (December 2010), 3.5 years old (June 2011) and four years old (December 2011). OPPA was planted entirely with oil palms; ground vegetation was few as compared to SF1 and SF2. SF1 remained undisturbed after logging activities whereas SF2 was disturbed forest due to minimal human activities.
About 100 g soil samples were collected using soil auger at 10 GPS-tagged sampling points (P1 to P10) from OPPA, SF1 and SF2. Each sampling points was at least 100 m apart. Four replications were taken from each sampling point and pooled as a composite sample to reduce any bias. All samples were collected at 0 to 15 cm depth after the removal of top vegetation with trowel. Soil samples were kept in sterile falcon tubes (Corning Inc.). New trowel was used and sterilised with 70% ethanol after each sampling sites. Soil samples were stored at -20°C for further use. This was to ensure that there was no microbial growth which could affect the result of this study. Layout of the study sites and sampling points is illustrated in Figure 1.
Soil Chemical Properties Analysis
Soil samples were air dried and 10 sampling points (P1 to P10) of each of the sampling sites were pooled together as one sample. The soil samples were analysed by a local agro-chemistry laboratory, RH Biotech Sdn. Bhd. in Sibu, Sarawak, for basic soil chemical analysis. Soil pH, total nitrogen (TN), total phosphorus (TP), exchangeable potassium (exch. K), and cation exchange capacity (CEC) were determined. Analysis of variance (ANOVA) was performed for the selected soil chemical properties with SAS – Statistical Analysis System (ver. 9.0, SAS Institute, Inc., Cary, NC, USA). Differences between sampling time were analysed using Tukey test at p≤0.05.
Characterisation of Soil Fungi Using DGGE
Extraction of DNA from soil samples was carried out using the PowerSoil DNA Isolation Kit (MO BIO) with a modified protocol. The procedures of PCR and DGGE were according to Vainio and Hantula (2000) with some modifications.
PCR Amplification of 18S rDNA Gene
PCR amplification of fungal 18S rDNA gene was performed using the fungi specific primers, GC-Fung and NS1 (May et al., 2001; White et al., 1990) under PCR condition as follows, initial denaturation at 94°C for 130 s, followed by 40 cycles of 94°C for 30 s, 55°C for 45 s and 72°C for 130 s and final extension at 72°C for 7 min (Cornejo et al., 2004). The sequences of the primers were as follows: primer NS1 (White et al., 1990), 5’-GTA GTC ATA TGC TTG TCT C-3’; primer GC-Fung (May et al., 2001), 5’-CGC CCG CCG CGC CCC GCG CCC GGC CCG CCG CCC CCG CCC CAT
TCC CCG TTA CCC GTT G-3’. The 40-nucleotide GC clamp was added to the 5’end of the reverse primer, to stabilise the melting behaviour of the PCR product for DGGE. Each PCR was carried out in a final volume of 25 µl, which contained 1 µl of DNA template, 0.25 U Taq polymerase, 1 x Reaction Buffer (Invitrogen), 2 mM MgCl2, 0.3 µM of each primers and 0.2 mM of dNTP. PCR products were analysed first by electrophoresis 1.0% agarose gel, PCR products loaded with EZ-Vision® (Amresco Inc., USA) and view under UV-illumination.
Denaturing Gradient Gel Electrophoresis (DGGE)
DGGE was performed in a D-Code system (Bio-Rad) with 16 cm x 16 cm x 1 mm thick of 7.5% polyacrylamide gels (arcylamide: bisacrylamide ratio 37.5:1) in 1x TAE buffer (40 mM Tris-acetate, 1 mM Na-EDTA; pH 8.0) and a 25%-40% gradient [100% corresponds to 7 M urea and 40% (v/v) deionised formamide]. Urea and formamide were used as the denaturing agents. Electrophoresis was carried out for 16 hr at temperature of 60°C and 55 V (Aguilera et al., 2006; Dı́ez et al., 2001). Marker was used as calibration among different lanes in each gel.
PCR of Excised Bands in TE Buffer
The targeted DNA was excised and eluted in TE buffer overnight. The entire eluted DNA was amplified with the same method as stated above, with primers without GC clamp. The PCR product was purified with GeneJETä (Fermentas). Later, the products were sent for DNA gene sequencing in commercial laboratory, First Base Laboratories Sdn. Bhd.
Analysis of DNA Sequences
Targeted gene sequence was edited and aligned using Bioedit software. After that the sequence was then compared with the gene sequences available in the NCBI database. List of blast results were compared to determine unique groups and common groups among assessment sites. Meanwhile, the number of phylum, class, order, family and genus were recorded as a way to monitor the composition of fungal community.
Analysis of DGGE Gel
Computer-assisted comparison of DGGE band profiles was analysed using Phoretix 1D gel analysis software (Total Lab Ltd., United Kingdom). Background noise was deducted by using the rolling ball algorithm with a radius of 50 pixels. First, automatic band detection was performed with a minimum slope of 75 to 100 and a noise reduction of 5, and peaks smaller than 2% of the maximum peak were eliminated. Bands positions were then compared with the DGGE marker to create a matrix containing the reference value (Rf) of each band.
The Rf of DGGE marker was set as 0.5 in every gel. The band matrix was then used to gain information for biodiversity indices and clustering (version 5; Primer-E Ltd, Plymouth, United Kingdom) to determine fungal communities in the OPPA and secondary forests, SF1 and SF2. Principal component analysis (PCA) was used to interpret each sampling because this analysis provides a way to reduce the complexity of our data to a lower dimensionality.
Biodiversity Indices
A diversity index is a measurement that shows how many different types (such as species) there are in the study area, and simultaneously takes into account how evenly the individuals are distributed among the community. The value of a diversity index increases as both the number of type increases and evenness increases. For a given number of types, the value of a diversity index is maximised when all types are equally abundant.
The Shannon diversity index, H’, a general diversity index, considers both species richness and evenness:
H’ = – Ʃ pi ln pi
where pi = ni/N, n is the total number of clones of the ith , and N the total number of samples.
The Simpson’s index of diversity was calculated as the following equation:
1-D = 1- {Ʃn (n-1)/N (N-1)}
where n equals to the total number of organisms of a particular species, N equals to the total number of organisms of all species. The value of D ranges between 0 and 1, thus 1 represents infinite diversity and 0, no diversity.
Soil Texture and Chemical Properties Analysis
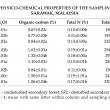
Both secondary forests of SF1 and SF2 were having the same soil texture, silty clay loam (SF1= 32% clay, 48% silt and 20% sand; SF2 = 32% clay, 49% silt and 19% sand). Soil texture in OPPA was loam (OPPA = 15% clay, 36% silt and 49% sand). These were determined with Texture Auto Lookup program based on US Department of Agriculture classification schemes (Teh and Rashid, 2003).
The selected soil chemical properties including soil pH, organic carbon (C), TN, TP, CEC, for each sampling period are summarised in Table 1.
Results showed that the conversion from secondary forest (SF1 and SF2) to OPPA had caused modifications to the physico-chemical properties of soil. Findings revealed that OPPA soils were more alkaline and with reduced organic C content. Soil pH would directly affects other soil chemical properties and influencing composition and diversity of soil fungi. The requirements of oil palm nutrients were estimated at around 110-156 kg ha–1 yr–1 for nitrogen (N), 64-98 kg ha–1 yr–1 for phosphorus (P), 10-21 kg ha–1 yr–1 for magnesium (Mg), 365-705 kg ha–1 yr–1 for potassium (K), in order to obtain optimum growth. The concentration of selected nutrients (TP, TN) in OPPA showed slight increase over the years (Table 1). This findings suggested that agricultural practices such as balance fertilisation that have been adopted could improve soil nutrients for the examined areas. However, the selected nutrients in SF1 showed slight decrease over the years. This may be due to leaching or natural soil erosion in the forest. Similarly, SF2 showed fluctuation in the soil nutrients, suggesting that minimal disturbances can have impacts to the soil.
CEC is important as an indicator to soil fertility because it shows the ability of the soil to provide calcium (Ca), Mg and K to the plants. From the results, OPPA gave lower CEC compared to SF2, and SF2 had lower CEC than SF1. This indicated that soil in OPPA was less fertile compared to SF1 and SF2, and SF2 was less fertile than SF1. High pH (less acidic) gave higher negative charge to the soil and hence, higher CEC. The soil pH of OPPA, SF1 and SF2 soils were strongly acidic because they ranged from pH 4.89 to pH 5.33. However, discrepancies were observed when SF1 showed higher pH (less acidic) yet yielded lower CEC. The soil chemical properties result demonstrated that soil type of SF1 was silty clay loam and thus, gave higher CEC, which was more fertile than OP soil. The selected soil chemical properties were subjected to a cluster analysis according to Euclidean distance analysis and summarised in Multidimensional Scaling (MDS) (Figure 2). Soil chemical properties of OPPA over years were in tight cluster, whereas soil chemical properties of SF1 and SF2 were in another cluster. This showed that soil chemical properties between OPPA and SF1, SF2 were significantly different. This was probably due to the effects of agricultural practices carried out in the oil palm plantation such as liming, pest and disease control and fertilising.
PCR Amplification of Partial 18S rDNA Gene
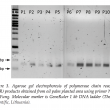
The 18S rDNA gene from all soil samples (n=90) were successfully amplified by primer pair NS1/GC-Fung (May et al., 2001; White et al., 1990) except one soil sample OP-3-P4. All the PCR products with the expected sizes of 370 bp were obtained. This indicated that no inhibition in the PCR products. No amplification in sample OP- 3-P4 might be due to the sample was taken from waterlogged area. Majority of soil fungi survive better in an aerobic conditions compared to anaerobic. This was explained by Ingham (1999) who stated that soil which is anaerobic due to waterlogged or compacted soils for significant periods, would lose their fungal component. Result of PCR products which successfully amplified for OPPA were shown in Figure 3.
DGGE Profiling of 18S rDNA Gene
The amplification of extracted soil DNA with fungal-specific primer pair, NS1/GC-fung (May et al., 2001) yielded products of 370 bp to be further investigated in DGGE. DGGE successfully separated all the DNA of same length into different bands. Results from the DGGE gels showed mixtures of bands with different intensities. This DGGE profile, or known as fingerprinting profile, contained the whole fungal community amplified from soil total DNA. Each discrete band is referred as operational taxonomic units (OTU) or discrete fungal population. Band intensity gives an idea to the frequency or abundance of individual fungal biomass. The number of bands corresponds to the number of members in the soil fungal communities (Duarte et al., 2012). Overall, high numbers of OTU were found in SF1. Whereas, lower OTU were recorded in both OP and SF2. Over hundreds of total number of bands were obtained in SF1 over the sampling time. Meanwhile, OPPA and SF2 recorded lesser total number of bands and most of the bands were separated at the middle part of the gels while fewer appeared at the top and bottom of the gels. Complex DGGE patterns were obtained, suggesting that complicated fungal communities existing in the soil.
Findings revealed that in agricultural land (OPPA), the fungal communities were much affected by their surroundings, including farm practices, pesticides and fertilisers applied to the palms. These activities directly affect physical, chemical and biological factors which resulted in reducing soil productivity. In forest area (SF1), the growth of soil fungi was between evenness and richness and that may refer to the natural habitat, humidity, shrubby and shading condition (Jena et al., 2015) and that matched with the finding revealed by OTU results in the current study. Whereby, OPPA in December 2010, there was less crop treatment being carried out while the oil palms were young and at new stage of growing. Bold dried land in OPPA was not a preferential habitat to the soil fungi. Therefore, soil fungal community was fewer.
Community Structure through Gene Sequence Analysis
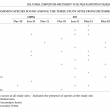
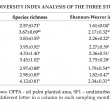
A total of 175 selected bands were excised from DGGE profile for further analysis using gene sequencing. Tables 2 and 3 show taxonomic order and common soil fungal species found in the study sites. Ten species were commonly found in both SF1 and SF2. Three common species were found in all assessment sites and they were A. polytricha, Clitocybula atroalba and S. cuboideum. Based on taxonomic order, five fungal species were from the Phylum Ascomycota (Aspergillus sp., A. ochraceus, S. cuboideum, Cochliobolus cynodontis, A. polytricha) and the rest were from the Phylum Basidiomycota.
The common species shared between the secondary forests SF1 and SF2 were Aspergillus sp., Antrodia xanthan, Cantharocybe gruberi, Cryptococcus podzolicus, H.cantharellus, Hygrocybe conica, Mycocali denudata, Ramaricium alboflavescens and Tricholoma subresplendens. Results showed that shifting of land from forest to oil palm plantation reduced the abundance of Aspergillus sp., A. ochraceus, Cantharocybe gruberi, C. podzolicus, H. cantharellus, I. conica, M. denudate, R. alboflavescens and T. subresplendens. Whereas, A. polytricha, C. atroalba, C. cynodontis and Rhizoctonia sp. were found to be resistant to deforestation. Sequencing of the excised bands gave the picture of composition of fungal community. Fungi from the phylum Ascomycota, genus Aspergillus were found in both SF1 and SF2. These fungi are common associates of decaying plant materials (Cannon and Kirk, 2007).
Basidiomycota were found in both SF1 and SF2, they belong to the family of Fomitopsidaceae, Tricholomataceae, Tremellaceae, Hygrophoraceae, Nidulariaceae and Gomphaceae (corresponding to the closest match A. xanthan, C. gruberi, C. podzolicus, H. cantharellus, H. conica, M. denudata, R. alboflavescens and T. subresplendens). Among these families, Fomitopsidaceae are the parasitic fungi causing brown rot on trees (Bolhassan et al., 2012). Hygrophoraceae are ectomycorrhizal or saprobic found in forests. Gomphaceae are widely spread in tropical climatic zones and formed ectomycorrhizal relationships with woody plants. Nidulariaceae are the saprobic on dead plant materials, especially on dead woods or grasses. Tremellaceae are often parasitic on other fungi. Tricholomataceae are ectomycorrhizal and occurs in broadleaved forests. All the ectomycorrhizals are dependent on the forest trees to survive, hence, they are absent from OPPA (Cannon and Kirk, 2007).
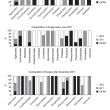
The composition of fungal orders increased over time as shown in the bar charts (Figure 4). SF1 and SF2 were having more diverse orders of fungi compared to OPPA. From the charts, it was obvious that fungi from the order Pleosporales were dominant in OPPA whereas the order Agaricales was dominant in SF1 and SF2. From the analysis of fungal orders, it was found that there were orders shared among OPPA, SF1 and SF2 and also there were orders uniquely exist at individual sites. Orders shared among OPPA, SF1 and SF2 were Agaricales, Polyporales, Pleosporales, Eurotiales and Cantharellales. They are terrestrial saprobes growing on fallen woods or grasses (Binder et al., 2013; Castellano et al., 2012).
Orders shared between OPPA and SF1 or SF2 are Auriculariales, Sordariales and Pezizales. These fungi are mostly saprobes on grasslands (Huhndorf et al., 2004). Auriculariales are jelly fungi (Sotome et al., 2014) whereas Pezizales are saprophytes and almost always occur on soil or dung (Harrington et al., 1999; Moore-Landecker, 1972). Orders shared between SF1 and SF2 are Phallales and Tremellales. Phallales grow on leaf litters in damp woodland (Grgurinovic and Simpson, 2001). Tremellales are parasites of other fungi (Pippola and Kotiranta, 2008). These were the common fungi found among OPPA, SF1 and SF2.
Unique orders in OPPA were Xylariales, Ascomycete, Microascales, Microthyrales and Sebacinales. Xylariales, Ascomycete, Microascales are saprobes found on terrestial environments (De Beer et al., 2013; Smith et al., 2003). Sebacinales are mycorrhizas (associated with plant roots for nutrition) (Weib et al., 2011). Unique orders in
SF1 are Aphyllophorales, Coniochaetales and Botrysphaeriales. Aphyllophorales are similar to Agaricales except they are not bearing gills (non- agarics) and they are root-rotting fungi in the soil (Rajchenberg, 1989). Botrysphaeriales are sac fungi and usually are saprobes (Adesemoye et al., 2014; Phillips et al., 2013). Coniochaetales are saprotrophs, leaf and root endophytes, and plant pathogens.
Unique orders in SF2 are Eurotiomycetes, Boletales, Corticiales, Onygenales, Russuales and Umbilicariales. Eurotiomycetes, Boletales, Corticiales are saprobes on wood and grasslands (Geiser et al., 2006). Onygenales are able to digest keratin and may be found on animals’ hooves in the forests (Johnston et al., 2015). Russulales could be found on hardwoods. Umbilicariales are lichenized fungi (Admčík et al., 2013; Miller et al., 2006). From the results above, it was observed that saprobes were among the most common fungi that were present in OPPA and secondary forests. The importance of fungi as saprobes in ecosystem processes is undeniable.
Ascomycota are predominantly terrestrial, some live in fresh or marine waters. They are majority of saprophytes on decaying plant debris or parasites on plants, insects and animals (Moore-Landecker, 1972). Ascomycota are important as decomposers (saprophytes). Together with other fungi, they break down large molecules in the environment and contribute to the nutrients cycling in the ecosystems. Some Ascomycetes form symbiotic relationship with other plants for survival strategy. They associate with green algae to form lichens; they live at the plant roots as mycorrhiza; and in plants to form endophytes. All these characteristics of Ascomycota make it a majority fungal composition in the oil palm planted areas.
Basically, Basidiomycetes follow the same roles as the other fungi; they are important decomposers, plant pathogens and symbionts with plants (mycorrhiza). Our results showed that Basidiomycota was the highest phylum in secondary forest. Our findings were in line with the reports of Moll et al. (2016), McGuire et al. (2015) and Dai et al. (2013). Basidiomycota are common wood decomposers or decomposers of leaf litters and animal dung. Some of them are mycorrhiza to the roots of forests trees (Moll et al., 2016). Hence, this explained why Basidiomycota are dominating secondary forest.
Forests floor were covered with decaying leaves litters. A study by Vorˇı´sˇkova´ and Baldrian (2013) showed that towards the later stage of degradation processes, fungi from the phylum Basidiomycota particularly the cellulolytic fungi would replace Ascomycota. This observation also supports our results of high abundance of Basidiomycota in secondary forests compared to oil palm plantation area. In addition, Basidiomycota are slow growing fungi and are sensitive to soil disturbance (McGuire et al., 2015). Therefore, Basidiomycetes are relatively low in oil palm plantation. Meanwhile, Ascomycota were recorded to be at higher abundance in stressful environments (Tripathi et al., 2016). Thus, this report is in line with our results.
PCA of DGGE Profiles
The binary matrix of DGGE bands were subjected to PCA (Figure 5). PCA is good in interpreting complex DGGE data, giving a quick and simple view to study the distribution of biodiversity. Findings revealed that the secondary forests (SF1 and SF2) shared similar DGGE banding patterns, yielding similar distribution when the scattered plot was produced. OPPA obviously produced unique DGGE banding pattern, thus, separated themselves to form one unique cluster. Groupings of SF1 and SF2 were found to be overlapped in June 2011, as this period was dry season. In short, SF1 and SF2 shared about similar DGGE banding pattern whereas OPPA was significantly different and was not significantly affected by dry or rainy seasons. This result suggested that soil fungal communities in OPPA were heterogeneous compared to the fungal community in SF1 and SF2. This observation matched previous results, where OPPA and SF1, SF2 were distinctive in their dominant fungal phylum.
Biodiversity of Soil Fungal Community
The total number of bands can be used for comparing fungal compositions (Duarte et al., 2012). Biodiversity index could be calculated by taking one band as one species to estimate the species richness and diversity. Binary matrix generated using CLIQS 1D Pro (Total Lab Ltd., United Kingdom) was used in the calculation of species richness. Shannon-Weaver index (H’) and Simpson (1-λ) were calculated by using software Primer-E (Plymouth). Table 4 shows the results of diversity index of the study sites. Species richness in OPPA, SF1 and SF2 at the beginning of the study were 2.57, 3.95 and 2.97, respectively. These indices increased to 3.83, 5.45 and 4.12 at the end of this study. In short, the species richness has increased significantly in every site from December 2010 to December 2011.
In Shannon-Weaver index, the increasing trend was observed in SF1 and SF2. OPPA recorded its Shannon-Weaver index as 1.6 (December 2010) to 2.26 (December 2011). However, there was no significant increment. SF1 and SF2 showed significant increment in Shannon-Weaver index. Shannon-Weaver index was higher in forest soils compared to OPPA. This was in line with a study by Grantina et al. (2011).
In Simpson index, there was no significant increment of soil fungal community in OPPA, SF2 and SF1. The diversity of soil fungal community showed increment over years in SF1 and SF2. SF1, SF2 and OPPA showed increment in species richness over the study. This showed that there was no significant disruption of composition of fungal community in SF1, SF2 and OPPA over the study period.
Apparently, our results showed that the composition and biodiversity of soil fungal communities in oil palm plantation have shifted in order to acclimatise with stressful environments as compared to secondary forest. Biodiversity of soil fungal communities in oil palm plantation reportedly lower than secondary forests and this suggested that plantation activities can cause destruction on the ecological components in the assessment sites. Hence, sustainable agricultural practices should be adopted in oil palm plantation to ensure good soil health and quality in order to conserve healthy ecosystems.
* Department of Animal Science and Fishery, Faculty of Agriculture and Food Sciences, Universiti Putra Malaysia Kampus Bintulu, 97008 Bintulu, Sarawak, Malaysia.
** Department of Agriculture Technology, Faculty of Agriculture, Universiti Putra Malaysia, 43400 UPM Serdang, Selangor, Malaysia.
‡ Department of Plant Protection, Faculty of Agriculture, Universiti Putra Malaysia, 43400 UPM Serdang, Selangor, Malaysia.
‡‡ Laboratory Sustainable Agronomy and Crop Protection (SACP), Institute of Plantation Studies (IKP), Universiti Putra Malaysia, 43400 UPM Serdang, Selangor, Malaysia.
E-mail: khairulmazmi@upm.edu.my